Rv0374c (-)
Current annotations:
TBCAP: (community-based annotations - see table at bottom of page )
TBDB: carbon-monoxide dehydrogenase small subunit
REFSEQ: carbon monoxyde dehydrogenase small subunit
PATRIC: Carbon monoxide dehydrogenase small chain (EC 1.2.99.2)
TUBERCULIST: Probable carbon monoxyde dehydrogenase (small chain)
NCBI: Probable carbon monoxyde dehydrogenase (small chain)
updated information (H37Rv4):
gene name: -
function:
reference:
Coordinates in H37Rv: 451800 - 452279
Gene length: 480 bp (with stop codon), 159 aa (without stop codon)
Operon:
Trans-membrane region:
Role: I.B.7 - Miscellaneous oxidoreductases and oxygenases
GO terms:
Reaction(s) (based on iSM810 metabolic model):
Gene Expression Profile Gene Modules Orthologs among selected mycobacteria Protein structure:
Search for Homologs in PDB Top 10 Homologs in PDB (as of Nov 2020): PDB aa ident species PDB title 1ZXI 59% Oligotropha carboxidovorans Reconstituted CO dehydrogenase from Oligotropha carboxidovorans 1N63 59% Oligotropha carboxidovorans Crystal Structure of the Cu,Mo-CO Dehydrogenase (CODH); Carbon monoxide reduced state 1N62 59% Oligotropha carboxidovorans Crystal Structure of the Mo,Cu-CO Dehydrogenase (CODH), n-butylisocyanide-bound state 1N61 59% Oligotropha carboxidovorans Crystal Structure of the Cu,Mo-CO Dehydrogenase (CODH); Dithionite reduced state 1N60 59% Oligotropha carboxidovorans Crystal Structure of the Cu,Mo-CO Dehydrogenase (CODH); Cyanide-inactivated Form 1N5W 59% Oligotropha carboxidovorans Crystal Structure of the Cu,Mo-CO Dehydrogenase (CODH); Oxidized form 4ZOH 57% Sulfolobus tokodaii (strain DSM 16993 / JCM 10545 / NBRC 100140 / 7) Crystal structure of glyceraldehyde oxidoreductase 1FFV 56% Hydrogenophaga pseudoflava CARBON MONOXIDE DEHYDROGENASE FROM HYDROGENOPHAGA PSEUDOFLAVA 1FFU 56% Hydrogenophaga pseudoflava CARBON MONOXIDE DEHYDROGENASE FROM HYDROGENOPHAGA PSEUDOFLAVA WHICH LACKS THE MO-PYRANOPTERIN MOIETY OF THE MOLYBDENUM COFACTOR 1T3Q 50% Pseudomonas putida Crystal structure of quinoline 2-Oxidoreductase from Pseudomonas Putida 86
Links to additional information on Rv0374c:
Amino Acid Sequence
MQVNMTVNGEPVTAEVEPRMLLVHFLRDQLRLTGTHWGCDTSNCGTCVVEVDGVPVKSCTMLAVMASGHSIRTVEGLAGPDGQLDPVQEGFMRCHGLQCG
FCTPGMLITARALLDRNPDPDEQTIREAISGQICRCTGYTTIVRSIQWAAAHQTVKAQS
(
Nucleotide sequence available on
KEGG )
Additional Information
MtbTnDB - interactive tool for exploring a database of published TnSeq datasets for Mtb
TnSeqCorr
Rv0374c/-,
gene len: 479 bp, num TA sites: 6
condition dataset call medium method notes
in-vitro DeJesus 2017 mBio non-essential 7H9 HMM fully saturated, 14 TnSeq libraries combined
in-vitro Sassetti 2003 Mol Micro growth-defect 7H9 TRASH essential if hybridization ratio<0.2
in-vivo (mice) Sassetti 2003 PNAS no data BL6 mice TRASH essential if hybridization ratio<0.4, min over 4 timepoints (1-8 weeks)
in-vitro (glycerol) Griffin 2011 PPath non-essential M9 minimal+glycerol Gumbel 2 replicates; Padj<0.05
in-vitro (cholesterol) Griffin 2011 PPath non-essential M9 minimal+cholesterol Gumbel 3 replicates; Padj<0.05
differentially essential in cholesterol Griffin 2011 PPath NO (LFC=-1.49) cholesterol vs glycerol resampling-SR YES if Padj<0.05, else not significant; LFC<0 means less insertions/more essential in cholesterol
in-vitro Smith 2022 eLife non-essential 7H9 HMM 6 replicates (raw data in Subramaniam 2017, PMID 31752678)
in-vivo (mice) Smith 2022 eLife non-essential BL6 mice HMM 6 replicates (raw data in Subramaniam 2017, PMID 31752678)
differentially essential in mice Smith 2022 eLife NO (LFC=1.109) in-vivo vs in-vitro ZINB YES if Padj<0.05, else not significant; LFC<0 means less insertions/more essential in mice
in-vitro (minimal) Minato 2019 mSys non-essential minimal medium HMM
in-vitro (YM rich medium) Minato 2019 mSys non-essential YM rich medium HMM 7H9 supplemented with ~20 metabolites (amino acids, cofactors)
differentially essential in YM rich medium Minato 2019 mSys NO (LFC=0.91) YM rich vs minimal medium resampling
Analysis of Positive Selection in Clinical Isolates
*new*
data from Culviner et al (2025) (55,259 Mtb clinical isolates)
overall pN/pS for Rv0374c: 1.030897969
lineage-specific pN/pS in L1: 1.090372852
lineage-specific pN/pS in L2: 0.726915234
lineage-specific pN/pS in L3: 0.908644043
lineage-specific pN/pS in L4: 1.138833867
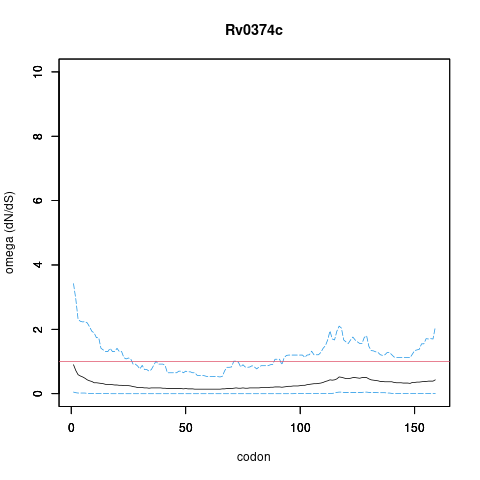
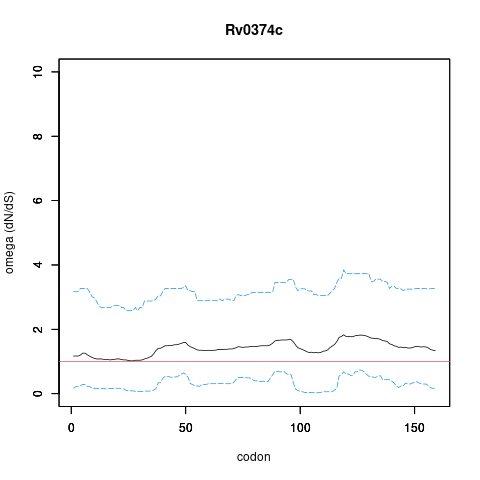
Analysis of dN/dS (omega) in two collections of Mtb clinical isolates using GenomegaMap (Window model) (see description of methods )
Moldova: 2,057 clinical isolates
global set: 5,195 clinical isolates from 15 other countries
In the omega plots, the black line shows the mean estimate of omega (dN/dS) at each codon, and the blue lines are the bounds for the 95% credible interval (95%CI, from MCMC sampling).
A gene is under significant positive selection if the lower-bound of the 95%CI of omega (lower blue line) exceeds 1.0 at any codon.
Moldova (2,057) global set (5,195)
under significant positive selection? NO NO
omega peak height (95%CI lower bound) 0.5 (0.05) 1.82 (0.74)
codons under selection
omega plots
genetic variants* link link
statistics at each codon link link
* example format for variants: "D27 (GAC): D27H (CAC,11)" means "Asp27 (native codon GAC) mutated to His (codon CAC) in 11 isolates"
TnSeq Data No data currently available.
No TnSeq data currently available for this Target.
RNASeq Data No data currently available.
No RNA-Seq data currently available for this Target.
Metabolomic Profiles No data currently available.
No Metabolomic data currently available for this Target.
Proteomic Data No data currently available.
No Proteomic data currently available for this Target.
Regulatory Relationships from Systems Biology
BioCyc
Gene interactions based on ChIPSeq and Transcription Factor Over-Expression (TFOE) (Systems Biology )
NOTE:
Green edges represent the connected genes being classified as differentially essential as a result of the middle gene being knocked out. These interactions are inferred based on RNASeq.
Interactions based on ChIPSeq data
RNA processing and modification
Energy production and conversion
Chromatin structure and dynamics
Amino acid transport and metabolism
Cell cycle control, cell division, chromosome partitioning
Carbohydrate transport and metabolism
Nucleotide transport and metabolism
Lipid transport and metabolism
Coenzyme transport and metabolism
Translation, ribosomal structure and biogenesis
Cell wall/membrane/envelope biogenesis
Replication, recombination and repair
Posttranslational modification, protein turnover, chaperones
Secondary metabolites biosynthesis, transport and catabolism
Inorganic ion transport and metabolism
General function prediction only
Intracellular trafficking, secretion, and vesicular transport
Signal transduction mechanisms
Differentially expressed as result of RNASeq in glycerol environment (Only top 20 genes shown sorted by log fold change with p_adj 0.05).
Conditionally essential as result of TNSeq (Only top 20 genes shown sorted by log fold change with p_adj 0.05).
Binds To:
No bindings to other targets were found.
Bound By:
No bindings from other targets were found.
Binds To:
No bindings to other targets were found.
Bound By:
Upregulates:
Does not upregulate other genes.
Upregulated by:
Not upregulated by other genes.
Downregulates:
Does not downregulate other genes.
Downregulated by:
Not downregulated by other genes.
Property Value Creator Evidence PMID Comment
Term TBRXN:CODH3 carbon monoxide dehydrogenase / acetyl-CoA synthase 2 - ISS njamshidi ISS 14660375|12486050 See PMID: 12486050, 14660375authors,GM. King Uptake of carbon monoxide and hydrogen at environmentally relevant concentrations by mycobacteria. Appl. Environ. Microbiol. 2003
Citation Growth of mycobacteria on carbon monoxide and methanol. authors,SW. Park,EH. Hwang,H. Park,JA. Kim,J. Heo,KH. Lee,T. Song,E. Kim,YT. Ro,SW. Kim,YM. Kim J. Bacteriol. 2003 njamshidi IPI 14660375|12486050 See PMID: 12486050, 14660375
Term TBRXN:CODH3 carbon monoxide dehydrogenase / acetyl-CoA synthase 2 - IPI njamshidi IPI 14660375|12486050 See PMID: 12486050, 14660375authors,SW. Park,EH. Hwang,H. Park,JA. Kim,J. Heo,KH. Lee,T. Song,E. Kim,YT. Ro,SW. Kim,YM. Kim Growth of mycobacteria on carbon monoxide and methanol. J. Bacteriol. 2003
Citation Uptake of carbon monoxide and hydrogen at environmentally relevant concentrations by mycobacteria. authors,GM. King Appl. Environ. Microbiol. 2003 njamshidi IPI 12486050|14660375 See PMID: 12486050, 14660375
Term TBRXN:CODH3 carbon monoxide dehydrogenase / acetyl-CoA synthase 2 - IPI njamshidi IPI 14660375|12486050 See PMID: 12486050, 14660375authors,GM. King Uptake of carbon monoxide and hydrogen at environmentally relevant concentrations by mycobacteria. Appl. Environ. Microbiol. 2003
Citation Growth of mycobacteria on carbon monoxide and methanol. authors,SW. Park,EH. Hwang,H. Park,JA. Kim,J. Heo,KH. Lee,T. Song,E. Kim,YT. Ro,SW. Kim,YM. Kim J. Bacteriol. 2003 njamshidi ISS 14660375|12486050 See PMID: 12486050, 14660375
Term TBRXN:CODH3 carbon monoxide dehydrogenase / acetyl-CoA synthase 2 - ISS njamshidi ISS 14660375|12486050 See PMID: 12486050, 14660375authors,SW. Park,EH. Hwang,H. Park,JA. Kim,J. Heo,KH. Lee,T. Song,E. Kim,YT. Ro,SW. Kim,YM. Kim Growth of mycobacteria on carbon monoxide and methanol. J. Bacteriol. 2003
Citation Uptake of carbon monoxide and hydrogen at environmentally relevant concentrations by mycobacteria. authors,GM. King Appl. Environ. Microbiol. 2003 njamshidi ISS 12486050|14660375 See PMID: 12486050, 14660375
Citation Heme oxygenase-1-derived carbon monoxide induces the Mycobacterium tuberculosis dormancy regulon. A. Kumar,JS. Deshane,DK. Crossman,S. Bolisetty,BS. Yan,I. Kramnik,A. Agarwal,AJ. Steyn J. Biol. Chem. 2008 akankshajain.21 IEP 18400743 Co-expression (Functional linkage)
Interaction Regulatory Rv3132c akankshajain.21 IEP Co-expression (Functional linkage)A. Kumar,JS. Deshane,DK. Crossman,S. Bolisetty,BS. Yan,I. Kramnik,A. Agarwal,AJ. Steyn Heme oxygenase-1-derived carbon monoxide induces the Mycobacterium tuberculosis dormancy regulon. J. Biol. Chem. 2008
Interaction Regulatory Rv2027c akankshajain.21 IEP Co-expression (Functional linkage)A. Kumar,JS. Deshane,DK. Crossman,S. Bolisetty,BS. Yan,I. Kramnik,A. Agarwal,AJ. Steyn Heme oxygenase-1-derived carbon monoxide induces the Mycobacterium tuberculosis dormancy regulon. J. Biol. Chem. 2008
Citation Heme oxygenase-1-derived carbon monoxide induces the Mycobacterium tuberculosis dormancy regulon. A. Kumar,JS. Deshane,DK. Crossman,S. Bolisetty,BS. Yan,I. Kramnik,A. Agarwal,AJ. Steyn J. Biol. Chem. 2008 prabhakarsmail IEP 18400743 Co-expression (Functional linkage)
Interaction Regulatory Rv3132c prabhakarsmail IEP Co-expression (Functional linkage)A. Kumar,JS. Deshane,DK. Crossman,S. Bolisetty,BS. Yan,I. Kramnik,A. Agarwal,AJ. Steyn Heme oxygenase-1-derived carbon monoxide induces the Mycobacterium tuberculosis dormancy regulon. J. Biol. Chem. 2008
Interaction Regulatory Rv2027c prabhakarsmail IEP Co-expression (Functional linkage)A. Kumar,JS. Deshane,DK. Crossman,S. Bolisetty,BS. Yan,I. Kramnik,A. Agarwal,AJ. Steyn Heme oxygenase-1-derived carbon monoxide induces the Mycobacterium tuberculosis dormancy regulon. J. Biol. Chem. 2008